Abstract
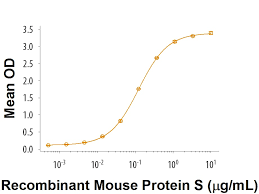
Homologous recombination (HR) is a high-precision DNA repair mechanism. Several HR genes are established cancer susceptibility genes with clinically actionable pathogenic variants (PVs). Classically, BRCA1 and BRCA2 germline PVs are associated with significant risks of breast and ovarian cancer. Patients with PV BRCA1 or BRCA2 show worse clinical outcomes but respond better to platinum-based chemotherapies and poly-ADP ribose polymerase inhibitors, a trait termed „BRCAness.“
With the advent of whole-exome sequencing and multigene panels, PVs in other HR genes are increasingly being identified among familial cancers. As such, several genes such as PALB2 are reclassified as cancer predisposition genes. But the evidence of cancer risks remains unclear for many others. In this review, we will discuss cancer recombinants predispositions and treatment implications beyond BRCA1 and BRCA2, with a focus on the 24 HR genes: 53BP1, ATM, ATR, ATRIP, BARD1, BLM, BRIP1, DMC1, MRE11A, NBN, PALB2, RAD50, RAD51, RAD51B, RAD51C, RAD51D, RIF1, RMI1, RMI2, RPA1, TOP3A, TOPBP1, XRCC2, and XRCC3.
Keywords
DNA double-strand break repair replication stress cogens-induced DNA damage model for cancer development DNA damage response genome instability homologous recombination inhibitors synthetic lethality RAD51BRCAness
Future directions
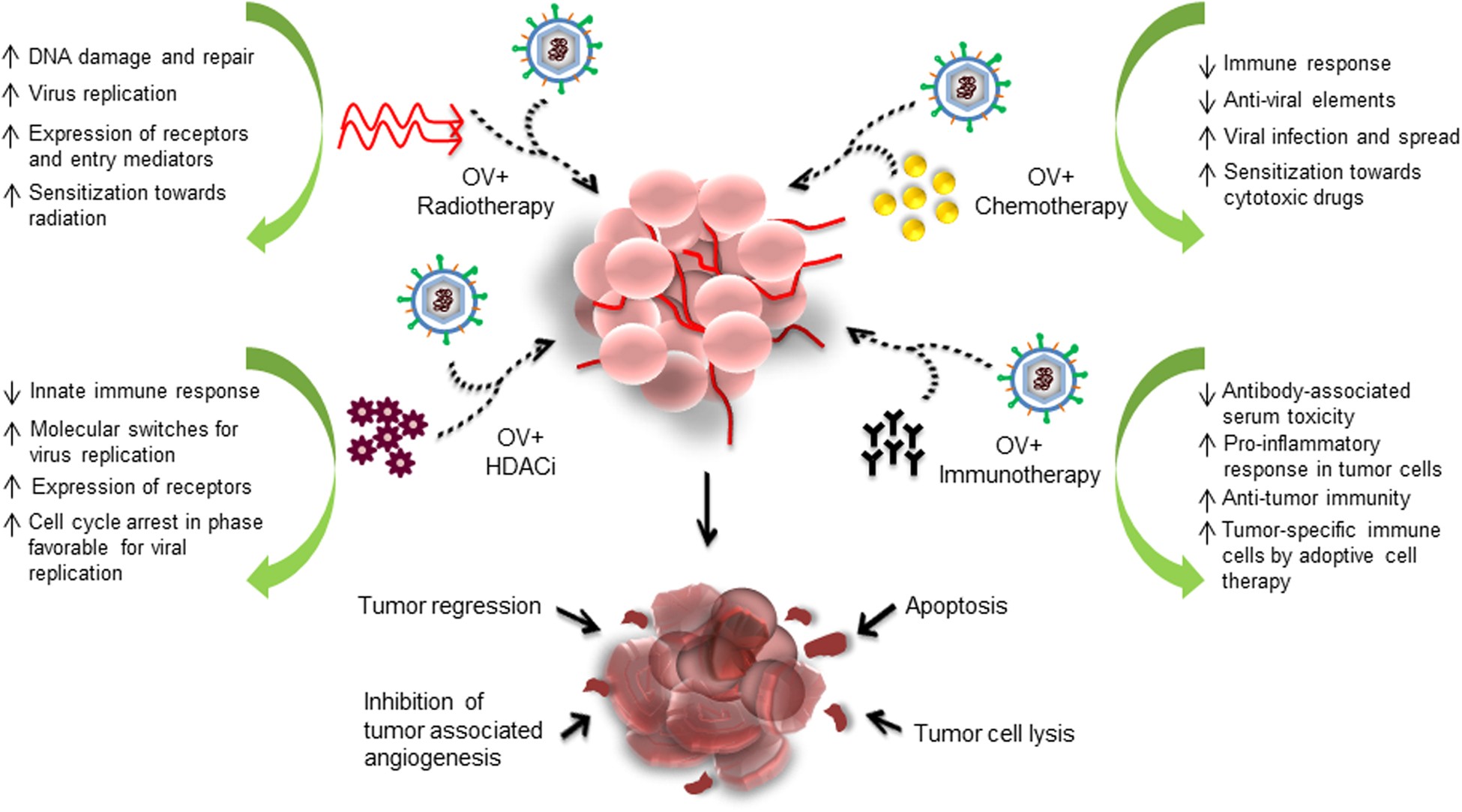
It is clear that HR has a great impact on cancer at various levels, especially in determining the efficacy of anticancer therapies. There is an urgent need to understand more about HR secondary pathways and how they differ from each other. Identification of proteins involved in these distinct HR pathways through large-scale RNA interference screens is highly important, as is further analysis of protein functions, to enable the development of better repair models. From Human Resources.
There is a need to determine the extent of HR defects in cancer and to characterize novel cancer-specific synthetic lethal interactions. Although such synthetic lethal interactions may not be strong enough to work in monotherapy, the increased therapeutic index could significantly improve chemotherapy combination treatments. Ultimately, we need better and more specific HR inhibitors to fully validate HR and its secondary pathways as relevant cancer targets.
Implications for Practice
This review provides a comprehensive reference for readers to quickly identify potential cancer-predisposing homologous recombination (HR) genes and generate research questions for genes with inconclusive evidence. This review also assesses the „BRCAness“ of each member of Human Resources. Physicians can refer to these discussions to identify potential candidates for future clinical trials.
Materials and methods
Homologous recombination genes were selected based on the authors‘ literature search. References for this review were identified through Google Scholar searches using the search terms “(gene name)”, “germline/somatic variants”, “prevalence”, “spectrum”, “multigene panel”, “ (type of cancer)”, “PARP inhibitors”, “platinum” and “BRCAness” with emphasis on articles published between 2016 and 2020.
Articles published in journals not indexed in PubMed were excluded. Abstracts published at established conferences were also reviewed. Only articles published in English were reviewed. The final reference list was generated based on originality and relevance to the broad scope of this review.
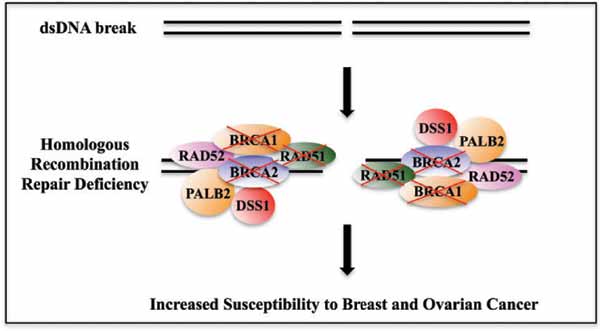
Definition and targeting of “BRCA quality”
Any defect in HR can lead to genome instability (ie, DNA deletions and chromosomal rearrangements) and trigger cell apoptosis. Tumours with inherited or sporadic HRD are more susceptible to DNA damage from platinum agents and PARPi, a trait known as „BRCAness.“ There are several ways to measure „BRCA capacity“: HRD, LST scores, tumour mutational signatures, and immunohistochemical detection of nuclear RAD5.
In addition to RAD51, we have introduced in this review several candidate genes whose abnormal expression may result in similar „BRCAness“ phenotypes. While some candidates are predictive of cancer risks and „BRCA capacity,“ the others are still of uncertain significance (see online Supplemental Data).
Conclusion
Historically, cancer treatment is largely based on the clinicopathological characteristics of the tumour and the affected organ. With the advent of genomic analysis and targeted therapies, the underlying genetic alteration of the tumour becomes relevant to its classification and treatment. Tumour HRD serves as a double-edged sword, correlating with a worse prognosis but better response to DNA-damaging agents. Although most inherited HRDs involve BRCA1 and BRCA2, PVs in non-BRCA HR genes contribute a significant portion of the „BRCA signature“ of the tumour. As our mechanistic understanding of HRD and synthetic lethality evolves, there will be more „drug“ HR targets for a broader spectrum of HR-deficient tumours.